There are currently are over 30,000 human diseases worldwide and effective treatments exist for only one third of them. Fortunately, there are millions of scientists around the world like yourself working relentlessly to better understand these diseases – and we are here to help them. This past year, Nicoya Lifesciences had the pleasure of helping innovative researchers publish more than 10 publications, by providing them with critical quantitative binding data. Dr. Xiaofeng Lu, a researcher at Sichuan University, uses OpenSPR’s surface plasmon resonance technology to provide her with the key binding data needed for her recent publication. The research publication “Targeted Delivery to Tumor-associated Pericytes via an Affibody with High Affinity for PDGFRβ Enhances the in vivo Antitumor Effects of Human TRAIL” uses SPR data generated with OpenSPR to study human tumor necrosis factor-related apoptosis using antibody targeting to deliver apoptotic drugs. Continue reading to learn how Dr. Lu used SPR data to discover an alternative strategy for tumor-targeted delivery of anticancer agents.
About the Publication
Previous studies have shown that human tumor necrosis factor-related apoptosis-inducing ligand (hTRAIL) exhibits in vitro cytotoxicity in a variety of tumor cells. However, hTRAIL did not exhibit an anticancer effect in clinical trials. This could be caused by hTRAIL being largely trapped by its decoy receptors, that are ubiquitously expressed on normal cells. Therefore, tumor-targeted delivery might improve the tumor uptake and enhance the antitumor effect of hTRAIL.
Tumor tissues derived both from patients with colon cancer and from mice with colorectal tumor xenografts are enriched with platelet-derived growth factor receptor β (PDGFRβ)-expressing pericytes. In this publication, a ZPDGFRβ affibody was distributed on tumor-associated PDGFRβ-positive pericytes. The ZPDGFRβ affibody mediated PDGFRβ-dependent binding of hTRAIL to pericytes was studied using surface plasmon resonance.
Why was OpenSPR instrumental for this research?
Surface plasmon resonance (SPR) was used to determine the binding between ZPDGFRβ affibody and PDGFRβ. The receptor fusion protein (PDGFRβ-Fc) was immobilized on an Nicoya’s carboxyl sensor chip, and the protein-protein experiment was run on Nicoya’s OpenSPR system. By using OpenSPR, Dr. Lu was able to determine the kinetic constants (association constant, dissociation constant and affinity) of the interaction between the affibody and hTRAIL complex to target the Fc receptor. More specifically, the ZPDGFRβ affibody was shown to bind to the receptor fusion protein (PDGFRβ-Fc) with a high affinity of 4.5 nM. In conclusion, the SPR data showed an increase tumor uptake of the apoptotic drug. With OpenSPR, Dr. Lu’s research lab was able to get SPR data from their own bench. This helped them accelerate their research and publish their discovery faster.
Why is SPR critical for publications? How does OpenSPR help?
SPR is a label-free technology which allows researchers to quantitatively analyze binding between two biomolecules. SPR technology allows us to determine the kon, koff and KD of interactions, providing deeper insight into binding events compared to other techniques that only give endpoint measurements, such as pull-down assays. SPR is necessary not only for publications but for the advancement of many fields of medicine and medical research as can be seen below with the significant increase in publications that rely on SPR data.
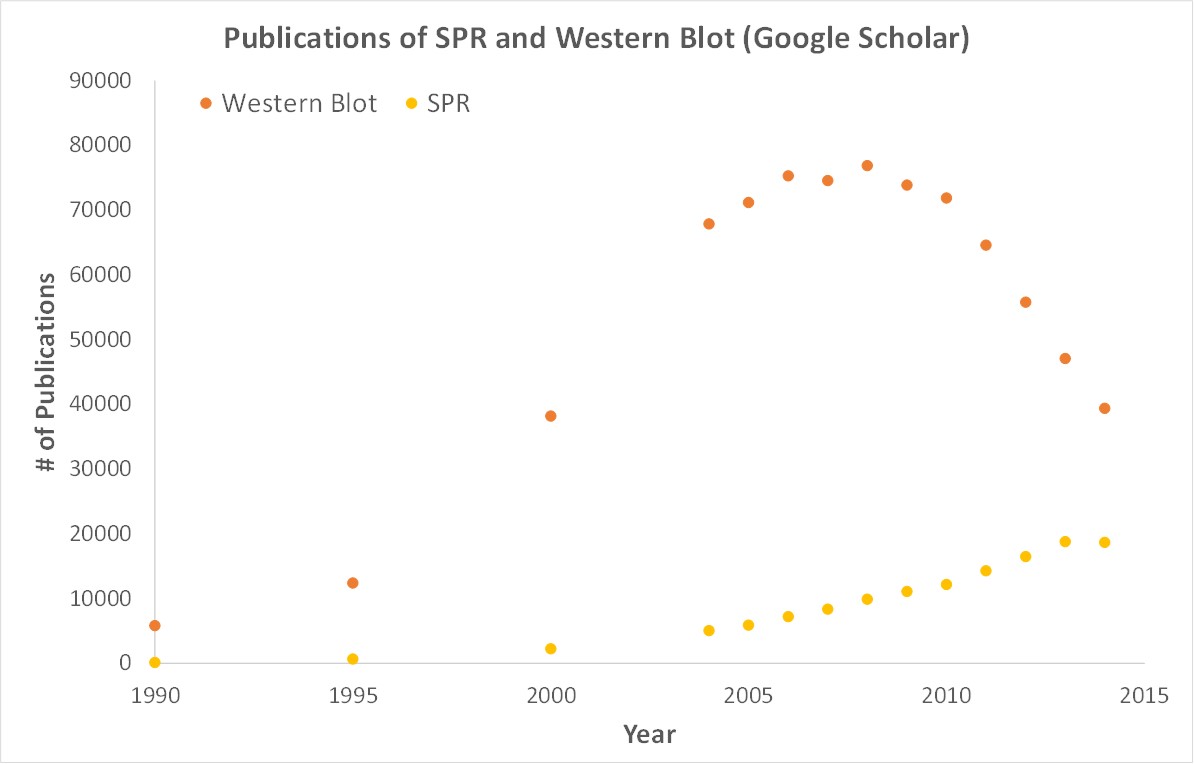
Scientific publications involving SPR have increased drastically over the years. SPR has become fundamental for publications while traditional techniques like Western Blots are becoming less important.
OpenSPR is a user-friendly and low maintenance benchtop SPR solution that is currently being used by hundreds of researchers. With access to SPR technology on your own lab bench you can get the high quality data you need to accelerate your research and publish faster.
