Your protein is precious. The more sample that you can save, the less time you need to spend on expressing and purifying proteins. This saves you money, and gives you more time to spend on other critical components of your research – like writing high-impact publications!
To obtain publication-quality binding kinetics data while only using small amounts of their protein samples, many researchers choose surface plasmon resonance (SPR). SPR provides quantitative kinetics data in real-time, and the OpenSPR™ offers all the benefits of SPR at a cost that is more accessible to researchers. However, to truly get the most out of your protein samples, SPR experts recommend using preconcentration screening with sensors that immobilize ligands via covalent coupling (such as Carboxyl and Amine Sensors). Using this method, the density of ligand immobilized to an SPR sensor can be increased substantially using only a small initial concentration of ligand. This means that an SPR experiment can be performed using even less of your protein sample. For even smaller sample sizes, check out Alto™ , which only uses 2 µL of analyte to conduct an entire binding experiment.
What is preconcentration?
Preconcentration is a covalent immobilization method that can be used to obtain a high ligand density at the sensor surface during immobilization. This is achieved by adjusting the pH of the buffer used for ligand immobilization (the immobilization buffer) to promote electrostatically driven accumulation of the protein on the sensor surface. For this reason, this technique only applies to sensor surface chemistries that exploit covalent coupling (such as Carboxyl and Amine Sensors) rather than a tag capturing method (such as NTA, GST, and Streptavidin sensors).
How preconcentration works
An activated Carboxyl Sensor surface is negatively charged at pH above 3 due to the acidic carboxyl (COOH) groups. The charge of the ligand (a protein in this case) is determined by the pH of the immobilization buffer and the isoelectric point (pI) of the ligand. By adjusting the pH of the immobilization buffer, it is possible to create opposite charges between the sensor surface and the ligand. This results in the ligand electrostatically accumulating on the surface of the sensor, which results in much higher densities of ligand immobilized than through simple surface collision. As a result, a local ligand concentration at the sensor surface could be greater than 100 mg/mL after starting with just 10 µg/mL of ligand.
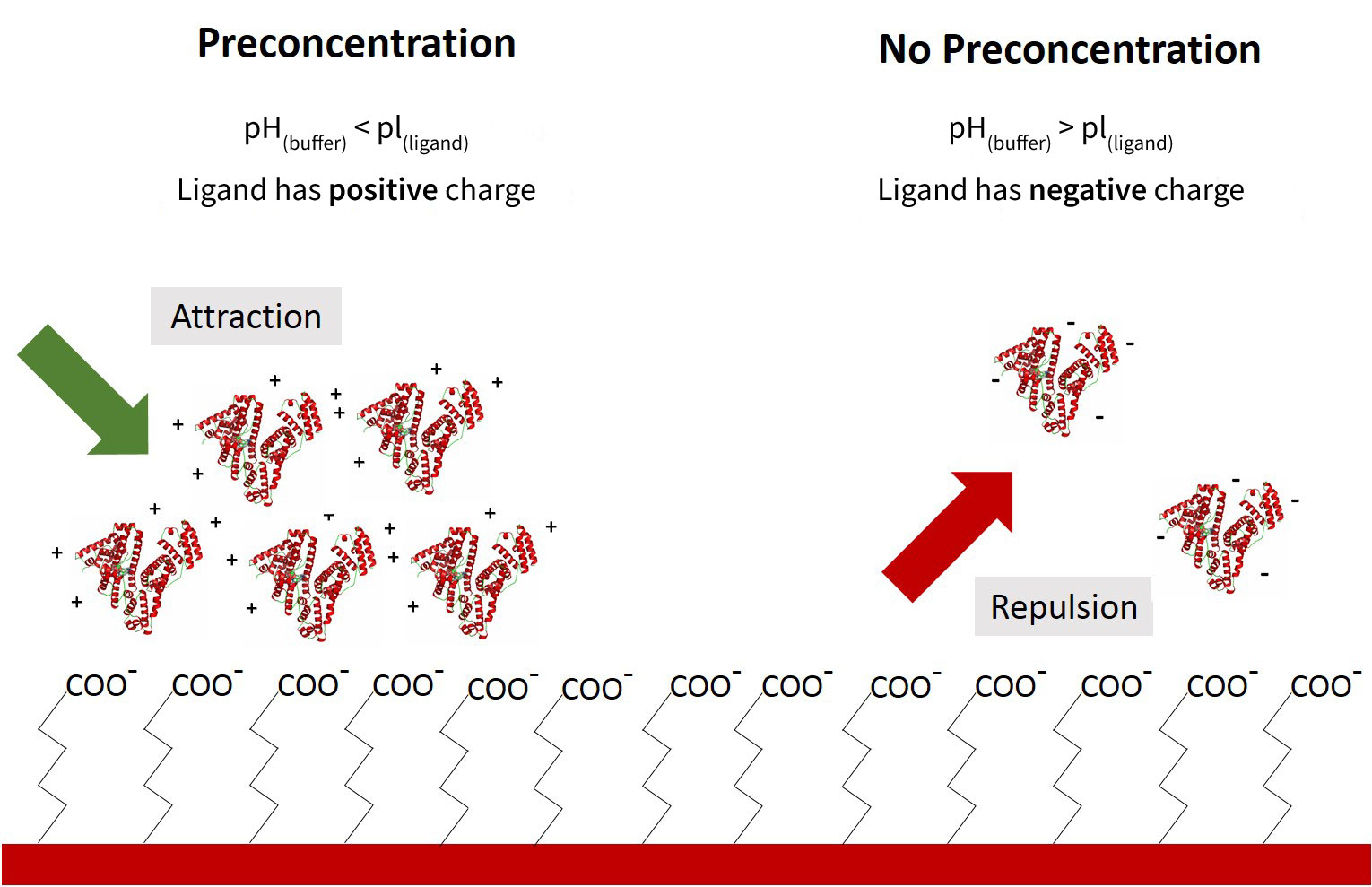
While a lower pH puts the ligand in a better state for pre-concentration, the actual covalent coupling occurs more efficiently at a higher pH. For this reason (and sometimes for ligand stability), a buffer pH that falls just below the pI of the ligand will theoretically provide the most optimal preconcentration conditions. Ultimately, the buffer conditions which yield the highest final immobilization density should be confirmed experimentally. While the Amine Coupling Kit used with OpenSPRTM Carboxyl Sensors is already optimized to effectively immobilize a wide variety of ligands, we recommend following the preconcentration protocol below to get the most out of your protein samples.
Optimization procedure
In order to determine the most optimal preconcentration conditions, we recommend testing ligand immobilization in immobilization buffers of varying pH. This can be done with a carboxyl senor that has not been activated with an Amine Coupling Kit so that the surface can be regenerated and used again several times. This way, preconcentration testing only requires one sensor chip. For convenience, the following steps can be followed on a 2-channel OpenSPR™. Note that this procedure uses an acetate buffer, but the same steps could be followed using the buffer more suitable for your system. More tips on buffer selection can be found here.
- Order an OpenSPR™ Immobilization Buffer Optimization Kit – this contains 10mM acetate buffers at pH 5.5, 5.0, 4.5 and 4.0 (20 mL of each) already prepared. These can also be prepared yourself.
- Dissolve the ligand at a low concentration (between 5-25 μg/ml) in each acetate buffer. Ensure that each buffer sample has the same concentration of ligand.
- Prepare a solution of 10 mM HCl to use as a regeneration solution.
- Load a standard Carboxyl Sensor into the OpenSPR™
- Wash with the regeneration solution
- Perform injection of the ligand in pH 5.5 acetate buffer through channels 1 and 2 at 10 uL/min
- Regenerate the sensor surface by injecting the regeneration solution at 100 μL/min.
- Repeat steps 6 and 7 using each buffer pH. We recommend testing each buffer pH twice – this would total 8 ligand injections
- Plot the resulting curves using TraceDrawer 1.8. The optimal immobilization buffer will have the highest pH that produced a large signal increase.
- The optimal immobilization buffer will have the highest pH that produced a large signal increase.
As an example, we performed a preconcentration experiment to determine the optimal buffer conditions to immobilize IgG. We followed the same procedure using 22.5 ug/mL igG solutions, and including a pH 6 acetate buffer in the analysis as well. Here are the results:
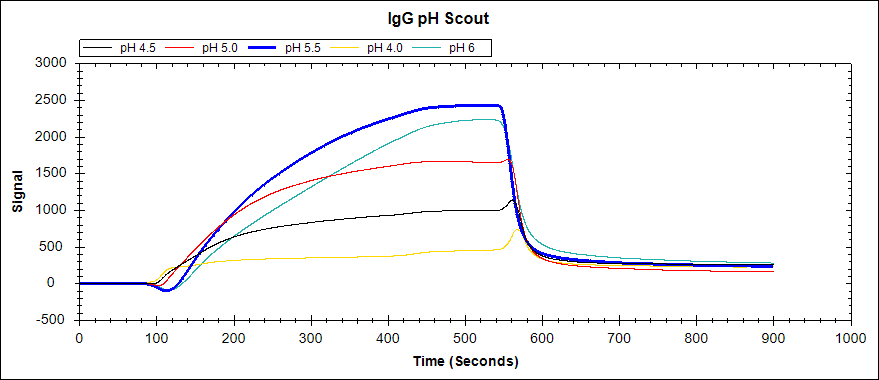
Figure 2. Immobilization of 22.5 ug/mL IgG to an unactivated Carboxyl Sensor in acetate buffers of varying concentrations.
As you can see, acetate buffers with pH 5.5 and pH 6 both resulted in a large signal increase. The higher pH of these two (pH 6) should then be used to immobilize igG in a binding experiment, as the Carboxyl Sensor will couple more efficiently at a higher pH.
Save your Protein Samples with OpenSPR™
At the end of the day, preconcentration really boils down to efficiency – getting the data you need while using as little of your precious protein as possible. That’s why many researchers use surface plasmon resonance (SPR) to study the binding of their proteins. SPR already requires small analyte volumes, and preconcentration helps you use even less of your ligand (for truly small sample sizes, check out our newest SPR system, Alto). SPR provides this quantitative data in real-time, without the use of labels. The OpenSPR™ provides all of these benefits at a price that allows researchers to use SPR on their own benchtop. With its intuitive design and our team of Customer Success Scientists helping you succeed, it has never been easier to introduce SPR into your experiment workflow.