You asked. We answered. Our team at Nicoya speaks with hundreds of researchers every week. We regularly hear that researchers want to perform kinetic analysis of analytes in crude samples. More specifically, when performing screening studies, the ability to quantify antibody-antigen binding in animal sera would be incredibly valuable in situations where antibody purification is not practical.
We are excited to share our latest application note: Binding Kinetics of an Antibody Serum Sample Using OpenSPR™. In this experiment, the OpenSPR™ was used to detect and measure the kinetics of anti-Mouse Immunoglobulin G (MIgG) in diluted antiserum. Using the OpenSPR™, our team was not only able to detect specific binding of an anti-mouse lgG in diluted serum, but was also able to obtain kinetic constants comparable to a positive control. In this blog, we explain the significance of serum sample analysis and then dive into this application note to highlight how the OpenSPR™ can be used to study analytes in complex samples.
Why is serum sample analysis important?
Antibody (Ab) screening is a critical step in the development of vaccines and other immunotherapies. Abs are evaluated for their ability to bind to specific targets. Surface plasmon resonance (SPR) is commonly used at this stage to obtain quantitative kinetic information about a panel of Abs so that researchers can make informed decisions when selecting candidates for further vaccine development. This is typically done with purified, monoclonal antibodies (mAbs) to ensure that impurities are not interfering and the observed binding is specific.
However, mAb production is a labour-intensive and extended process. It starts with polyclonal antibodies from animal sera, followed by a ranking/selection process. The selected Abs then undergo subsequent cloning or sequencing. Only a few (or none!) of the millions of Abs produced in an animal may be chosen for this process.
Therefore, it is important to only select the Abs with the desired binding properties (such as affinity), which may require screening of whole (not purified) anti sera for binding candidates. Fortunately, as our latest application note shows, the OpenSPR™ can be used to perform kinetic analysis of serum samples in situations where it is not practical to fully purify the analytes.
Kinetic analysis of an antibody serum sample with the OpenSPR™
In our latest application note, Mouse Immunoglobulin G (MIgG) was immobilized to an OpenSPR™ Carboxyl Sensor following standard amine coupling procedure. In addition to the blocking solution used in the Amine Coupling Kit, Bovine Serum Albumin (BSA) was used as a blocking agent in the reference channel to help prevent non-specific binding.
Human Immunoglobulin G (HIgG) was then analyzed as a negative control. The absence of binding with the negative control confirms that the serum analyte binding observed is specific (Figure 1). Next, purified Polyclonal Goat Anti-mouse (IgG) was analyzed as a positive control; Finally, goat serum containing anti-MIgG was analyzed (Figure 2).
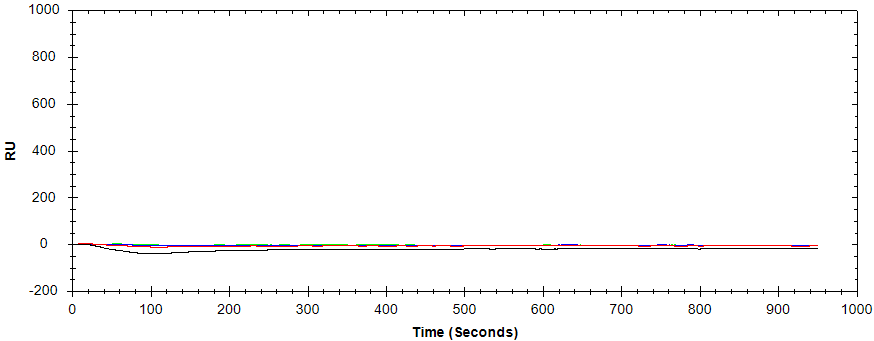
Figure 1: Human IgG (negative control) injected over the immobilized ligand at concentrations of 400 nM, 80 nM, 16 nM, and 3.2 nM.
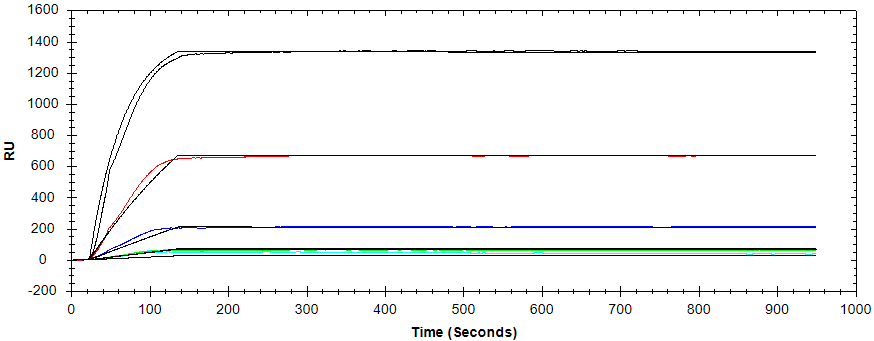
Figure 2: Binding of anti-MIgG at 600 nM, 120 nM, 24 nM, 4.8 nM, and 0.96 nM to the immobilized MIgG. The solid black lines represent the one-to-one kinetic model fits.
The positive and negative control experiments allude to the accuracy of the serum sample experiment. Also, the lack of interference from impurities is shown by the comparability of the kinetic data for the purified antibody versus the whole serum sample:
Table 1: Kinetic values measured using OpenSPR™ data with the TraceDrawer™ analysis software. The first kinetic values represent the analyte (goat anti-mouse IgG, whole Antiserum), followed by the positive control (purified polyclonal goat anti-mouse IgG).
| Whole Serum MIgG | Purified MIgG | |
| kon [1/M*s] | 3.77e4 (±3.39e0) | 3.36e4 (±1.07e0) |
| koff [1/s] | 5.47e-6 (±3.82e-6) | 4.94e-6 (±8.19e-6) |
| KD | 1.45e-10 (±1.01e-10) | 1.47e-10 (±2.44e-10) |
Therefore, this study demonstrates that the OpenSPR™ can be used to precisely detect and measure the binding kinetics of an analyte in a diluted complex medium. With the user-friendly design and affordable price of the OpenSPR™, it has never been easier to measure serum samples and a diverse range of other biomolecular interactions on your own lab bench.
