In order for drugs to fight diseases, an efficient delivery method must be found. This necessitates an important area of study to ensure that drugs can effectively combat diseases. A subsection of this ever-expanding field is nanomedicine; as it’s name implies, nanomedicine straddles the fields of medicine and nanotechnology, in an effort to increase drug specificity, bioavailability and biodistribution. A hallmark of this drug delivery approach includes employment of nanoparticles and oftentimes, the immune system views these as foreign inclusions and neutralizes these vehicles. In this respect, it is important to construct candidate nanotherapeutics in a more physiologically-relevant manner. Importantly, cell membrane camouflaging is a novel drug delivery method that has demonstrated promising clinical performance.
In this new publication, Dr. Chong Li et al. used OpenSPR’s localized surface plasmon resonance technology to provide them with the key binding data needed for their latest discovery. The publication titled, “Oriented Assembly of Cell-Mimicking Nanoparticles via a Molecular Affinity Strategy for Targeted Drug Delivery” uses binding kinetics data generated from the OpenSPR to explore the binding behaviours of the P4.2 peptide to the band 3 protein to probe the efficacy of cell membrane cloaking as a nanotherapeutic delivery system.
[et_pb_section bb_built=”1″ admin_label=”section”][et_pb_row admin_label=”row” background_position=”top_left” background_repeat=”repeat” background_size=”initial”][et_pb_column type=”4_4″][et_pb_text admin_label=”Paragraph 1″ background_layout=”light” text_orientation=”justified” text_font_size=”16″ text_line_height=”1.9em” use_border_color=”off” custom_margin=”-25px|||” custom_padding=”||10px|” background_position=”top_left” background_repeat=”repeat” background_size=”initial” _builder_version=”3.0.96″]
About the Publication
[/et_pb_text][/et_pb_column][/et_pb_row][et_pb_row background_position=”top_left” background_repeat=”repeat” background_size=”initial”][et_pb_column type=”4_4″][et_pb_text admin_label=”Paragraph 2″ background_layout=”light” text_font_size=”16″ text_line_height=”1.9em” use_border_color=”off” custom_margin=”-20px|||” background_position=”top_left” background_repeat=”repeat” background_size=”initial” _builder_version=”3.0.96″ /][et_pb_text admin_label=”Paragraph 2″ background_layout=”light” text_font_size=”16″ text_line_height=”1.9em” use_border_color=”off” custom_margin=”-20px|||” background_position=”top_left” background_repeat=”repeat” background_size=”initial” _builder_version=”3.0.96″ header_line_height=”5px”]
Cell membrane cloaking is an emerging drug delivery strategy in which a synthetic nanoparticle can be coated with an active cell membrane which resembles the composition of the targeted cell type. This method of drug delivery provides the advantages of self-recognition, immune evasion, increased bioavailability and may be applied to many clinical applications including cancer and deadly infections.
Importantly, band 3, found in the cellular membrane of Red blood cells (RBCs), is a transmembrane protein responsible for chloride exchange across the erythrocyte membrane. Notably, band 3 binds a cytoplasmic protein known as protein 4.2 with fairly high affinity and this interaction is thought to play a role in RBC shape regulation. Since the affinity between these two proteins of interest is quite high, the authors posture that it’s possible to exploit the interaction in the context of nanotherapeutic construction. However, adequate preparation of the delivery vehicle is required to realize the advantages of immune evasion, bioavailability and self-recognition. Using Candida albicans as a model infection, the authors prepared a peptide of protein 4.2 to specifically recognize the cytoplasmic domain of band 3 to construct a nanodelivery system with high efficacy.
In this publication, the investigators employed a number of in vivo and in vitro techniques to design, construct and evaluate a novel nanotherapeutic delivery system. A peptide of protein 4.2 specific to the cytoplasmic domain of band 3 was engineered via molecular modeling and binding constants were computationally derived. Subsequently, RBC membrane-coated liposomes (RBC-LIP) were prepared as delivery vehicles and the correct peptide orientation was confirmed with transmission electron microscopy and fluorescence colocalization assays. In vivo imaging in mouse lungs and receptor occupancy assays with flow cytometry were conducted to confirm binding of the delivery constructs. Notably, these techniques only provided relative binding information and surface plasmon resonance was recruited to obtain the on rates, off rates and equilibrium constants of band 3 and peptide 4.2.
[et_pb_section bb_built=”1″ admin_label=”section”][et_pb_row admin_label=”row” background_position=”top_left” background_repeat=”repeat” background_size=”initial”][et_pb_column type=”4_4″][et_pb_text admin_label=”Paragraph 1″ background_layout=”light” text_orientation=”justified” text_font_size=”16″ text_line_height=”1.9em” use_border_color=”off” custom_margin=”-25px|||” custom_padding=”||10px|” background_position=”top_left” background_repeat=”repeat” background_size=”initial” _builder_version=”3.0.96″]
Why was OpenSPR instrumental for this research?
[/et_pb_text][/et_pb_column][/et_pb_row][et_pb_row background_position=”top_left” background_repeat=”repeat” background_size=”initial”][et_pb_column type=”4_4″][et_pb_text admin_label=”Paragraph 2″ background_layout=”light” text_font_size=”16″ text_line_height=”1.9em” use_border_color=”off” custom_margin=”-20px|||” background_position=”top_left” background_repeat=”repeat” background_size=”initial” _builder_version=”3.0.96″ /][et_pb_text admin_label=”Paragraph 2″ background_layout=”light” text_font_size=”16″ text_line_height=”1.9em” use_border_color=”off” custom_margin=”-20px|||” background_position=”top_left” background_repeat=”repeat” background_size=”initial” _builder_version=”3.0.96″ header_line_height=”5px”]
Surface plasmon resonance (SPR) was used to directly measure binding constants between the cytoplasmic domain of band 3 and peptide 4.2; importantly, full-length protein 4.2 was also employed in a solution competition binding assay. In the direct binding assay, the His-tagged band 3 protein was immobilized on an NTA sensor chip and triplicate concentrations of peptide 4.2 were injected at a flow rate of 20 uL/min. Importantly, the measured KD value for band 3 and peptide 4.2 was 2.69 nM, indicating a solid interaction between the ligand and analyte and thus, specificity of peptide 4.2 for band 3. In the solution competition assay, GST-tagged, full-length recombinant protein 4.2 was immobilized on the sensor and increasing concentrations (0, 40, 80, 160 and 200 nM) of peptide 4.2 and static concentrations of band 3 (20 nM) were co-injected. Notably, a reduced response was observed when the 40nM concentration of peptide 4.2 was co-injected with band 3, indicating higher affinity of peptide 4.2 to band 3 than full-length protein 4.2 and confirming specificity. Upon completion of OpenSPR experiments, the authors posture that a “molecular affinity” strategy should be employed during the construction of nanotherapeutic delivery systems.
In this respect, OpenSPR binding data allowed researchers to evaluate a nanodelivery construction method suitable for clinical and industrial purposes. By using OpenSPR, the researchers were able to get SPR data from their own bench, to help them accelerate their research and publish their discovery faster.
[/et_pb_text][/et_pb_column][/et_pb_row][et_pb_row background_position=”top_left” background_repeat=”repeat” background_size=”initial”][et_pb_column type=”4_4″][et_pb_text admin_label=”Paragraph 2″ background_layout=”light” text_font_size=”16″ text_line_height=”1.9em” use_border_color=”off” custom_margin=”-20px|||” background_position=”top_left” background_repeat=”repeat” background_size=”initial” _builder_version=”3.0.96″ /][et_pb_text admin_label=”Paragraph 2″ background_layout=”light” text_font_size=”16″ text_line_height=”1.9em” use_border_color=”off” custom_margin=”-20px|||” background_position=”top_left” background_repeat=”repeat” background_size=”initial” _builder_version=”3.0.96″ header_line_height=”5px”]
Why is SPR critical for publications? How does OpenSPR help?
[/et_pb_text][/et_pb_column][/et_pb_row][et_pb_row background_position=”top_left” background_repeat=”repeat” background_size=”initial”][et_pb_column type=”4_4″][et_pb_text admin_label=”Paragraph 2″ background_layout=”light” text_font_size=”16″ text_line_height=”1.9em” use_border_color=”off” custom_margin=”-20px|||” background_position=”top_left” background_repeat=”repeat” background_size=”initial” _builder_version=”3.0.96″ /][et_pb_text admin_label=”Paragraph 2″ background_layout=”light” text_font_size=”16″ text_line_height=”1.9em” use_border_color=”off” custom_margin=”-20px|||” background_position=”top_left” background_repeat=”repeat” background_size=”initial” _builder_version=”3.0.96″ header_line_height=”5px”]
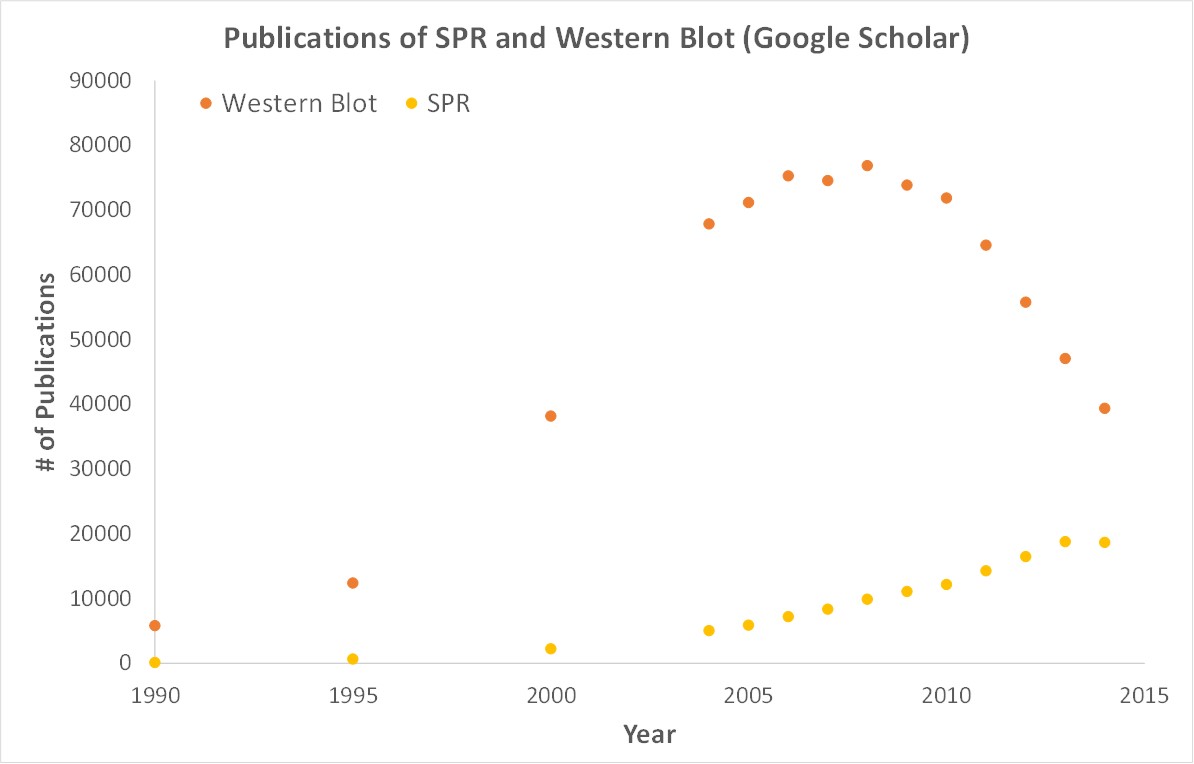
SPR is a label-free technology which allows researchers to quantitatively analyze binding between two biomolecules. SPR technology allows us to determine the kon, koff and KD of interactions, providing deeper insight into binding events compared to other techniques that only give endpoint measurements, such as pull-down assays. SPR is necessary not only for publications but for the advancement of many fields of medicine and medical research as can be seen below with the significant increase in publications that rely on SPR data.
[et_pb_section bb_built=”1″ admin_label=”section”][et_pb_row admin_label=”row” background_position=”top_left” background_repeat=”repeat” background_size=”initial”][et_pb_column type=”4_4″][et_pb_text admin_label=”Paragraph 1″ background_layout=”light” text_orientation=”justified” text_font_size=”16″ text_line_height=”1.9em” use_border_color=”off” custom_margin=”-25px|||” custom_padding=”||10px|” background_position=”top_left” background_repeat=”repeat” background_size=”initial” _builder_version=”3.0.96″]
[/et_pb_text][/et_pb_column][/et_pb_row][et_pb_row background_position=”top_left” background_repeat=”repeat” background_size=”initial”][et_pb_column type=”4_4″][et_pb_text admin_label=”Paragraph 2″ background_layout=”light” text_font_size=”16″ text_line_height=”1.9em” use_border_color=”off” custom_margin=”-20px|||” background_position=”top_left” background_repeat=”repeat” background_size=”initial” _builder_version=”3.0.96″ /][et_pb_text admin_label=”Paragraph 2″ background_layout=”light” text_font_size=”16″ text_line_height=”1.9em” use_border_color=”off” custom_margin=”-20px|||” background_position=”top_left” background_repeat=”repeat” background_size=”initial” _builder_version=”3.0.96″ header_line_height=”5px”]
OpenSPR is a user-friendly and low maintenance benchtop SPR solution that is currently being used by hundreds of researchers. With access to SPR technology on your own lab bench you can get the high-quality data you need to accelerate your research and publish faster.